Simulations
Research Areas
Confirmation of Bioinformatics Predictions of the Structural Domains in Honeybee Silk
Polymers 2018, 10, 776.
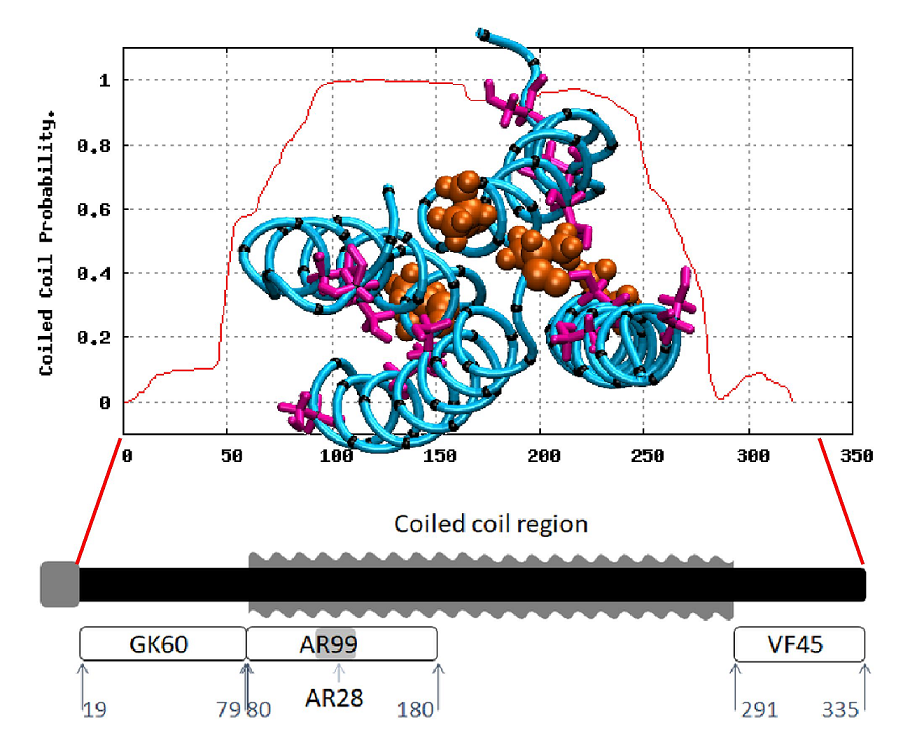
Abstract: Honeybee larvae produce a silk made up of proteins in predominantly a coiled coil molecular structure. These proteins can be produced in recombinant systems, making them desirable templates for the design of advanced materials. However, the atomic level structure of these proteins is proving difficult to determine: firstly, because coiled coils are difficult to crystalize; and secondly, fibrous proteins crystalize as fibres rather than as discrete protein units. In this study, we synthesised peptides from the central structural domain, as well as the N- and C-terminal domains, of the honeybee silk. We used circular dichroism spectroscopy, infrared spectroscopy, and molecular dynamics to investigate the folding behaviour of the central domain peptides. We found that they folded as predicted by bioinformatics analysis, giving the protein engineer confidence in bioinformatics predictions to guide the design of new functionality into these protein templates. These results, along with the infrared structural analysis of the N- and C-terminal domain peptides and the comparison of peptide film properties with those of the full-length AmelF3 protein, provided significant insight into the structural elements required for honeybee silk protein to form into stable materials.
Keywords: coiled coil; protein secondary structure; bioinformatics protein folding prediction; protein design; protein engineering; protein materials; infrared spectroscopy; circular dichroism spectroscopy; molecular dynamics; cast film solubility
Full Text
Improving the Description of Interactions Between Ca2+ and Protein Carboxylate Groups, Including γ-Carboxyglutamic Acid: Revised CHARMM22* Parameters
RSC Advances 2015, 83(5). 67820-67828
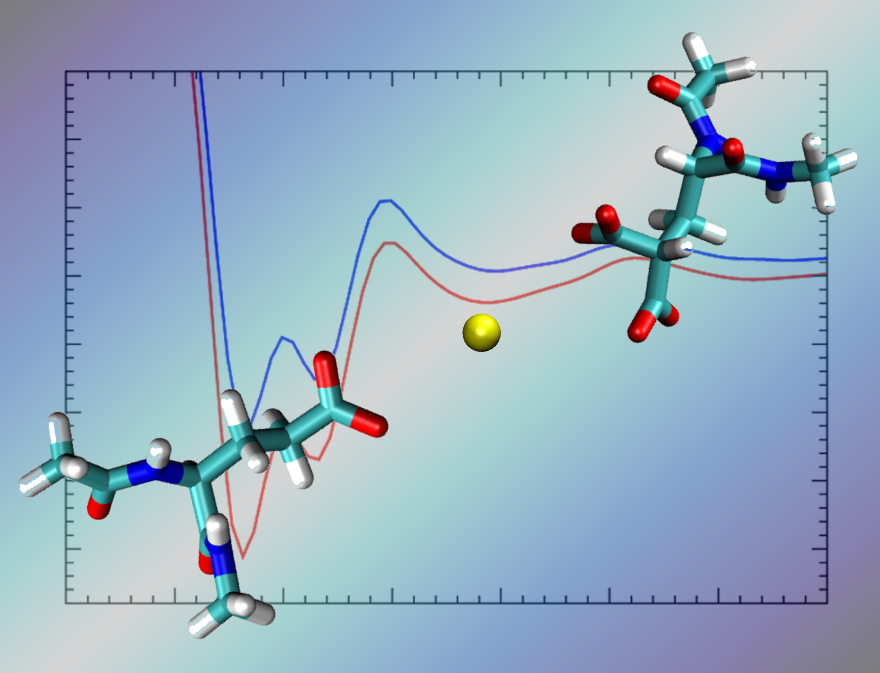
Abstract: A reliable description of ion pair interactions for biological systems, particularly those involving polyatomic ions such as carboxylate and divalent ions such as Ca2+, using biomolecular force-fields is essential for making useful predictions for a range of protein functions. In particular, the interaction of divalent ions with the double carboxylate group present in γ-carboxyglutamic acid (Gla), relevant to the function of many proteins, is relatively understudied using biomolecular force-fields. Using force-field based metadynamics simulations to predict the free energy of binding between Ca2+ and the carboxylate group in liquid water, we show that a widely-used biomolecular force-field, CHARMM22*, substantially over-estimates the binding strength between Ca2+ and the side-chains of both glutamic acid (Glu) and Gla, compared with experimental data obtained for the analogous systems of aqueous calcium–acetate and calcium–malonate. To correct for this, we propose and test a range of modifications to the σ value of the heteroatomic Lennard–Jones interaction between Ca2+ and the oxygen of the carboxylate group. Our revised parameter set can recover the same three association modes of this aqueous ion pair as the standard parameter set, and yields free energies of binding for the carboxylate–Ca2+ interaction in good agreement with experimental data. The revised parameter set recovers other structural properties of the ion pair in agreement with the standard CHARMM22* parameter set.
Full Text